The Trouble with Evolution
With millions of pieces of evidence, the theory of evolution is as factual as, say, Newton’s laws of motion! Yet, how it’s taught in schools requires a reexamination before it can achieve its intended goals of education. Some items need attention before introducing the subject to the interested parties.
Lamarck’s theory
The theory that says evolution is the adaptation of organisms to their environment has only historical relevance. It is not how evolution works. People are stuck to Lamarck’s theory, partly because it was taught before Darwin’s and also because it fits our fantasy of conversion and purpose. We will address these two terms soon. To repeat: individual organisms don’t evolve or pass their aspirations to offspring through genes.
Metaphors taken literally
We already know that nature doesn’t select anybody. Also, physical strength and superiority have nothing to do with the survival probability of a species. Yet, we carry the burden of natural selection and survival of the fittest in their literal meaning. These terms are strictly metaphors to communicate, perhaps wrong choices from people who lived a hundred years ago!
Another common feature in science communication is to say genes want to copy and spread. It creates a false notion of purpose in listeners’ minds. Again, a gene has no brain to decide anything, unlike humans, who are involved in designing artefacts for their use. You know, this purpose is not that purpose!
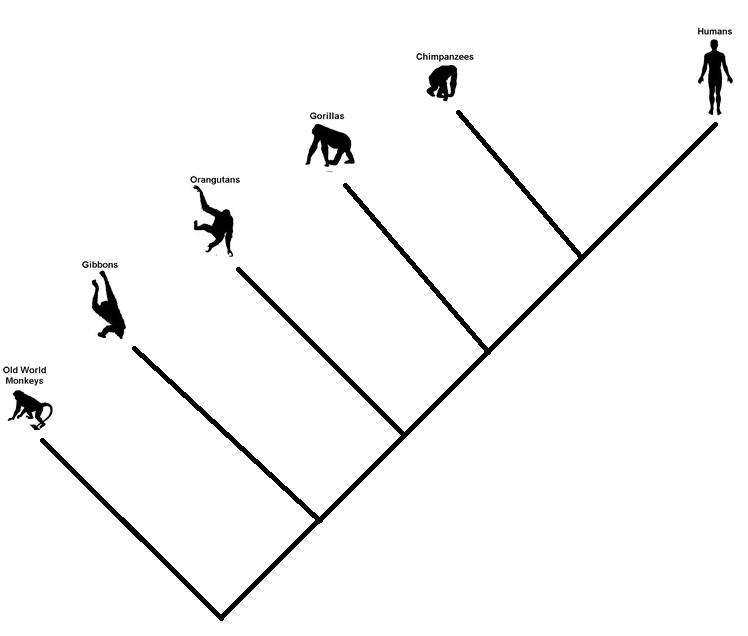
It goes in branches
This one came from a cartoonist – the money to the man. It is not a conversion process that works linearly. Once a species passes the baton, it doesn’t exit the scene. Evolution steps are random branching processes. So monkeys may survive, and so do apes or the great apes. Some may perish as well.
In summary
The features we see in today’s organisms are not part of any plans for perfection but simply a collection of clues about our past.
The Trouble with Evolution Read More »